Monica Civera and Laura Belvisi
Department of Chemistry, Università di Milano
Cyclic peptides and peptidomimetics are emerging as promising therapeutics for the modulation of protein-protein interactions because of their favorable properties, in terms of pharmacokinetic, bioavailability, metabolic stability and target specificity[1]. For instance, cyclization, possibly combined with other conformational constraints, can significantly decrease the entropic penalty to obtain an active conformation, enhancing the binding affinity to a specific receptor.
However, the computational study of cyclic systems is hampered by the critical problem to calculate their three-dimensional structures, both in the unbound and the bound state. Indeed, due to the constrained nature of cyclic peptides, most docking algorithms are unable to accomplish the concerted motions required for rigorous macrocycle conformation sampling. As a result, the use of poorly representative input structures strongly affects the predictability of molecular docking results. Thus, the exhaustive and reliable study of cyclopeptide conformations is increasingly regarded as an essential step prior to docking calculations and the use of multiple conformers has become the standard for the docking of cyclic systems[2].
Nevertheless, the comprehensive conformational sampling of cyclic peptides and peptidomimetics is not trivial: in these molecules, the investigation of equilibrium conformations is limited by the difficulties to cross high free-energy barriers and to perform the concerted rotation of several dihedrals required for the transition from one conformation to another. Moreover, the force fields implemented in molecular modeling softwares could be not appropriate for the structure determination of highly constrained cyclic peptides containing non-standard amino acids or peptidomimetic scaffolds.
In this workshop, the challenging issues affecting the computational study of cyclic peptides and peptidomimetics and some approaches for their solution will be discussed in the context of integrin ligands, where several cyclic peptides reached advanced phases of clinical investigation[3].
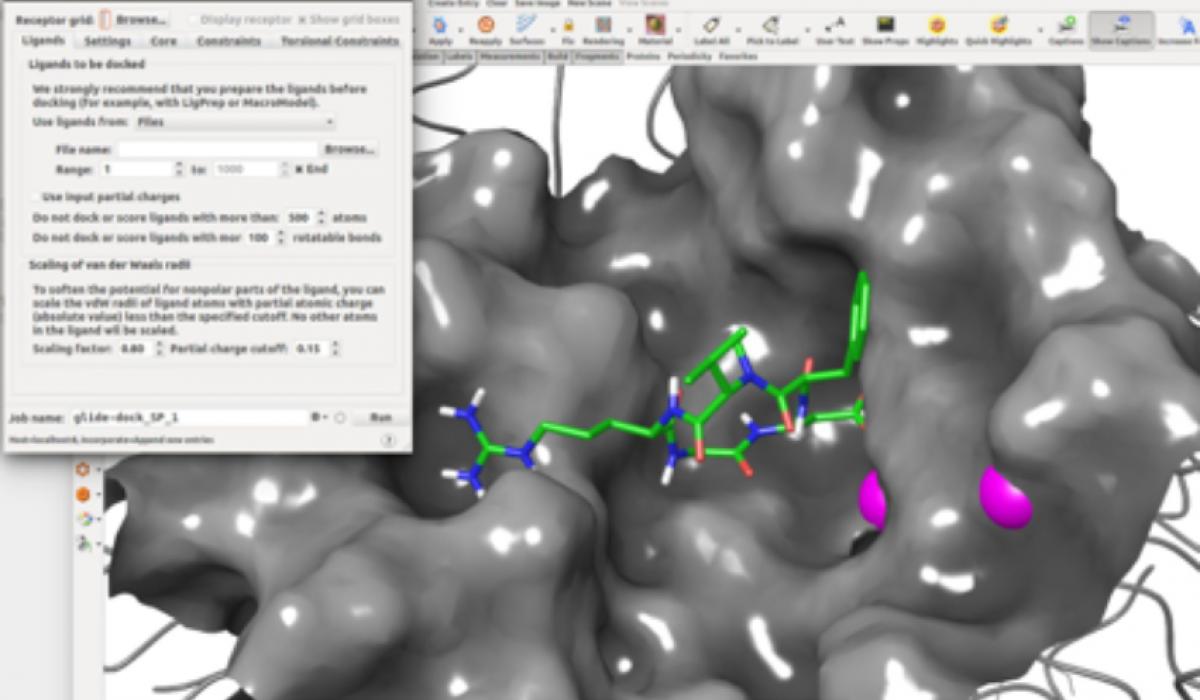
The presentation will also include an introduction to some tools from Schrodinger[4] that have been successfully used in the study of cyclic peptidomimetic integrin ligands[5].
REFERENCES:
[1] L. Nevola, E. Giralt, Chem. Commun. 2015, 51, 3302-3315. [2] S. E. Allen, N. V. Dokholyan, A. A. Bowers, ACS Chem. Biol. 2016, 11, 10−24. [3] C. Mas-Moruno, R. Fraioli, F. Rechenmacher, S. Neubauer, T. G. Kapp, H. Kessler, Angew. Chem. Int. Ed. 2016, 55, 7048 – 7068, and references therein. [4]https://www.schrodinger.com [5] M. Marchini, M. Mingozzi, R. Colombo, I. Guzzetti, L. Belvisi, F. Vasile, D. Potenza, U. Piarulli, D. Arosio, C. Gennari, Chem. Eur. J. 2012, 18, 6195−6207; M. Mingozzi, A. Dal Corso, M. Marchini, I. Guzzetti, M. Civera, U. Piarulli, D. Arosio, L. Belvisi, D. Potenza, L. Pignataro, C. Gennari, Chem. Eur. J. 2013, 19, 3563−3567.
PRESENTED IN:
